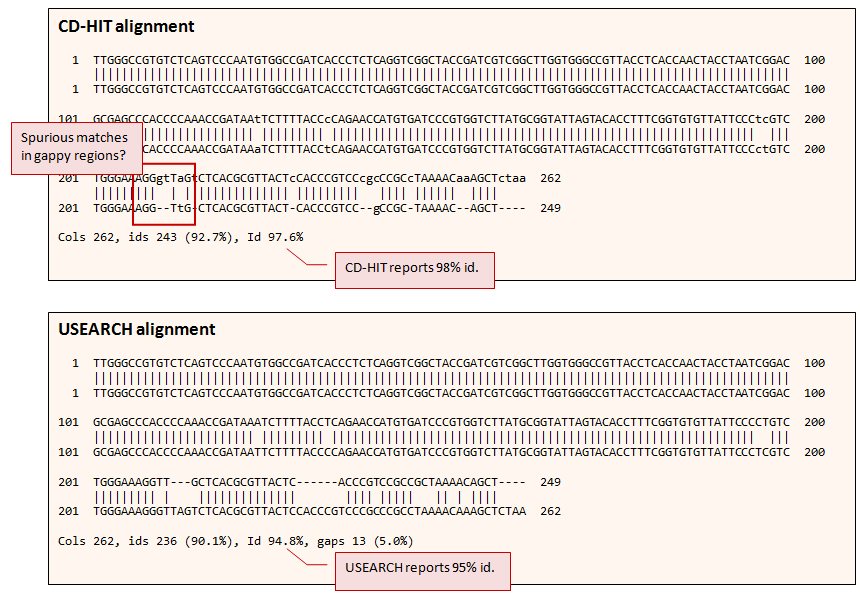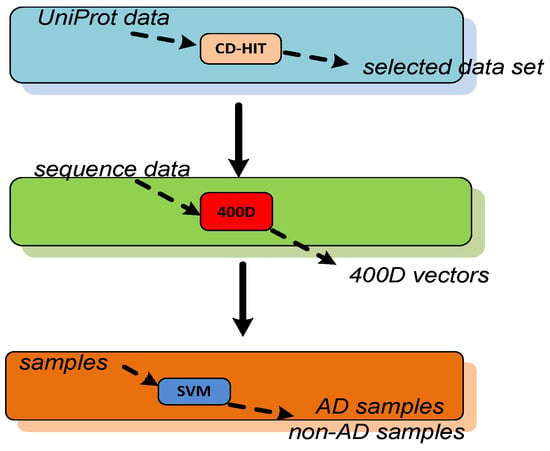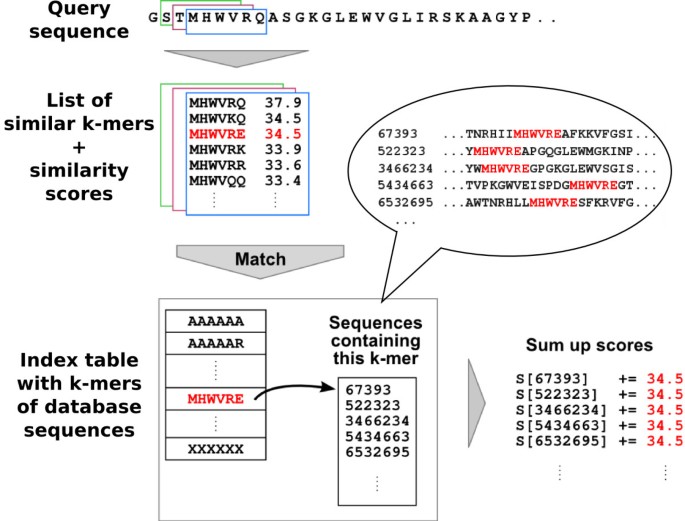
kClust: fast and sensitive clustering of large protein sequence databases | BMC Bioinformatics | Full Text

SUMOgo: Prediction of sumoylation sites on lysines by motif screening models and the effects of various post-translational modifications | Scientific Reports
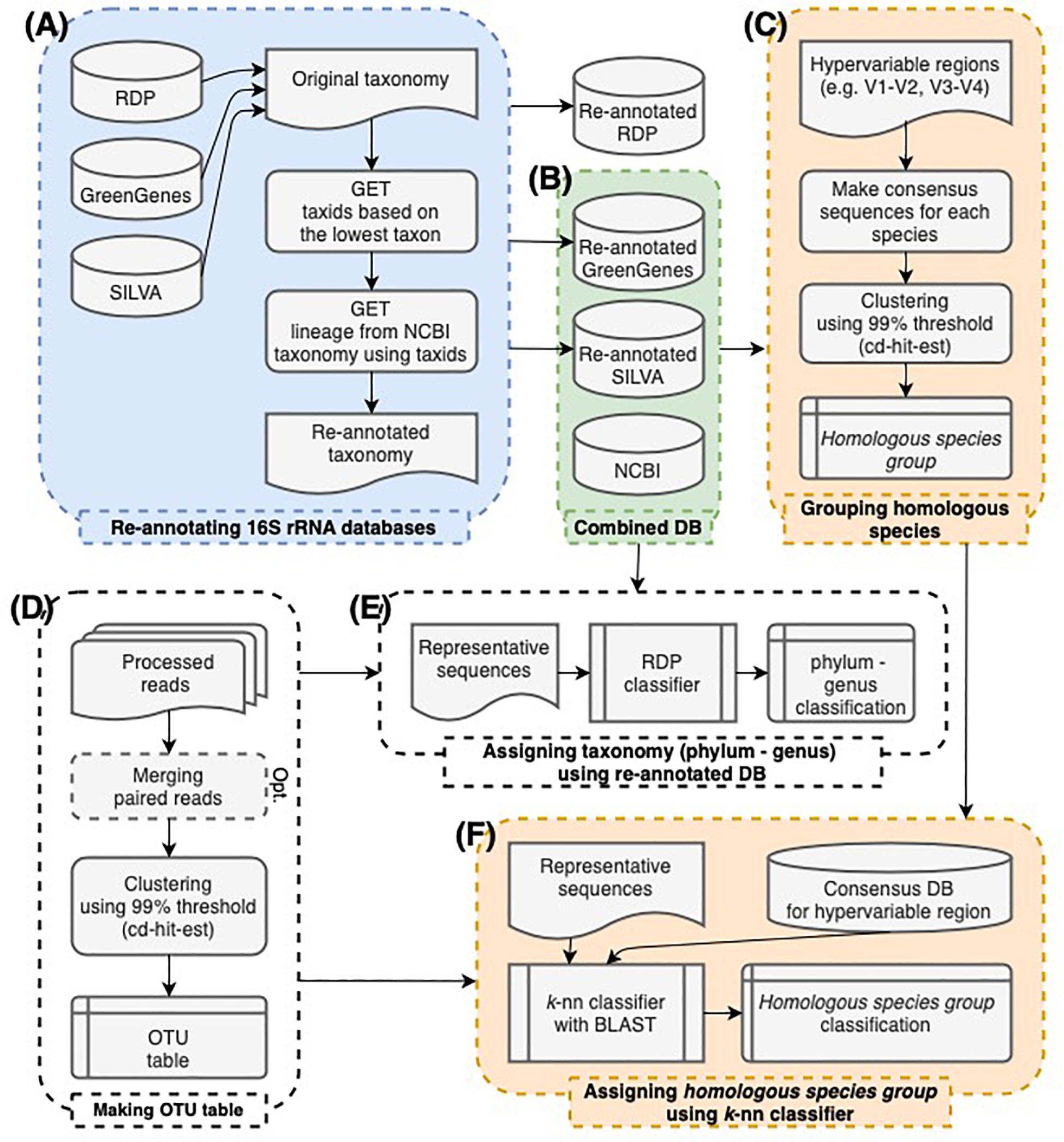
Frontiers | Data-Driven Modeling for Species-Level Taxonomic Assignment From 16S rRNA: Application to Human Microbiomes
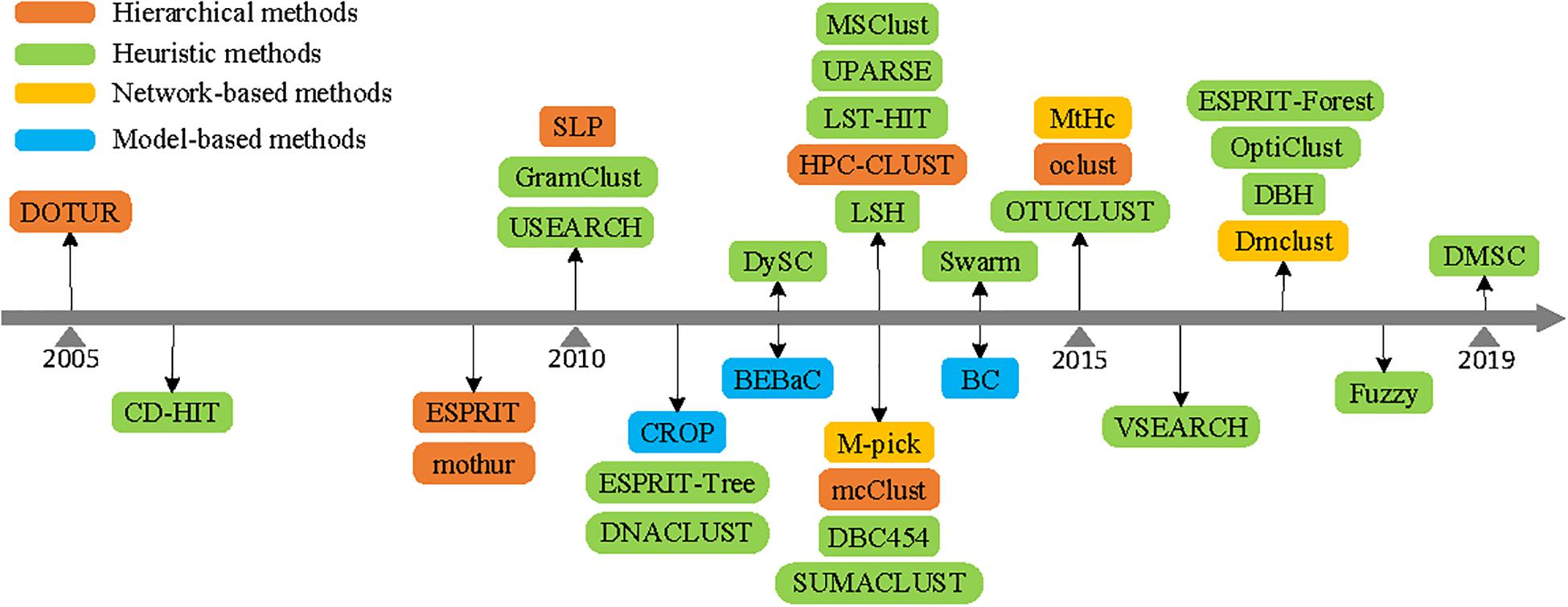
Frontiers | Comparison of Methods for Picking the Operational Taxonomic Units From Amplicon Sequences




/20039968/3380596/cd_cover.jpeg)