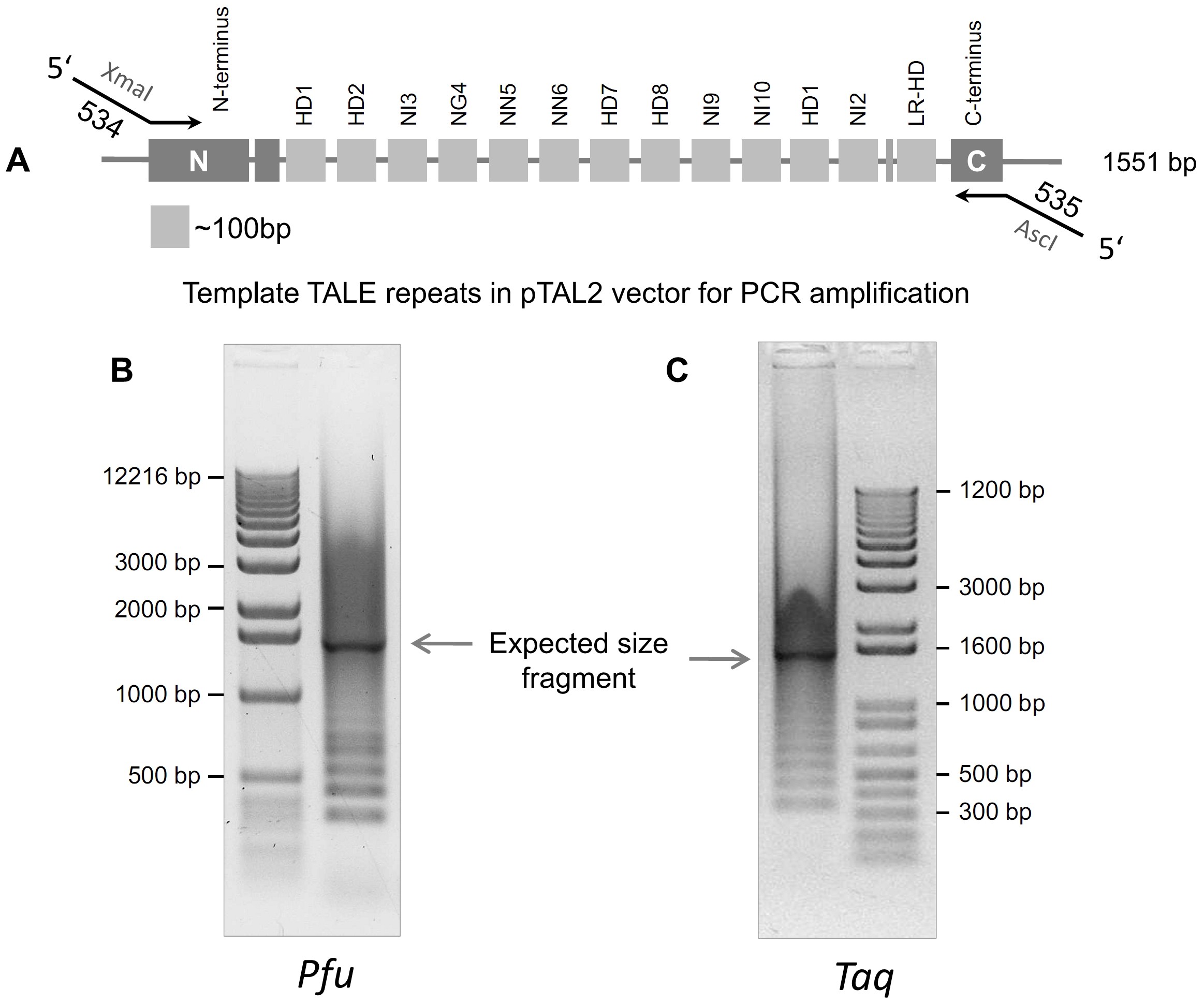
Primer evaluation and development of a droplet digital PCR protocol targeting amoA genes for the quantification of Comammox in lakes | Scientific Reports
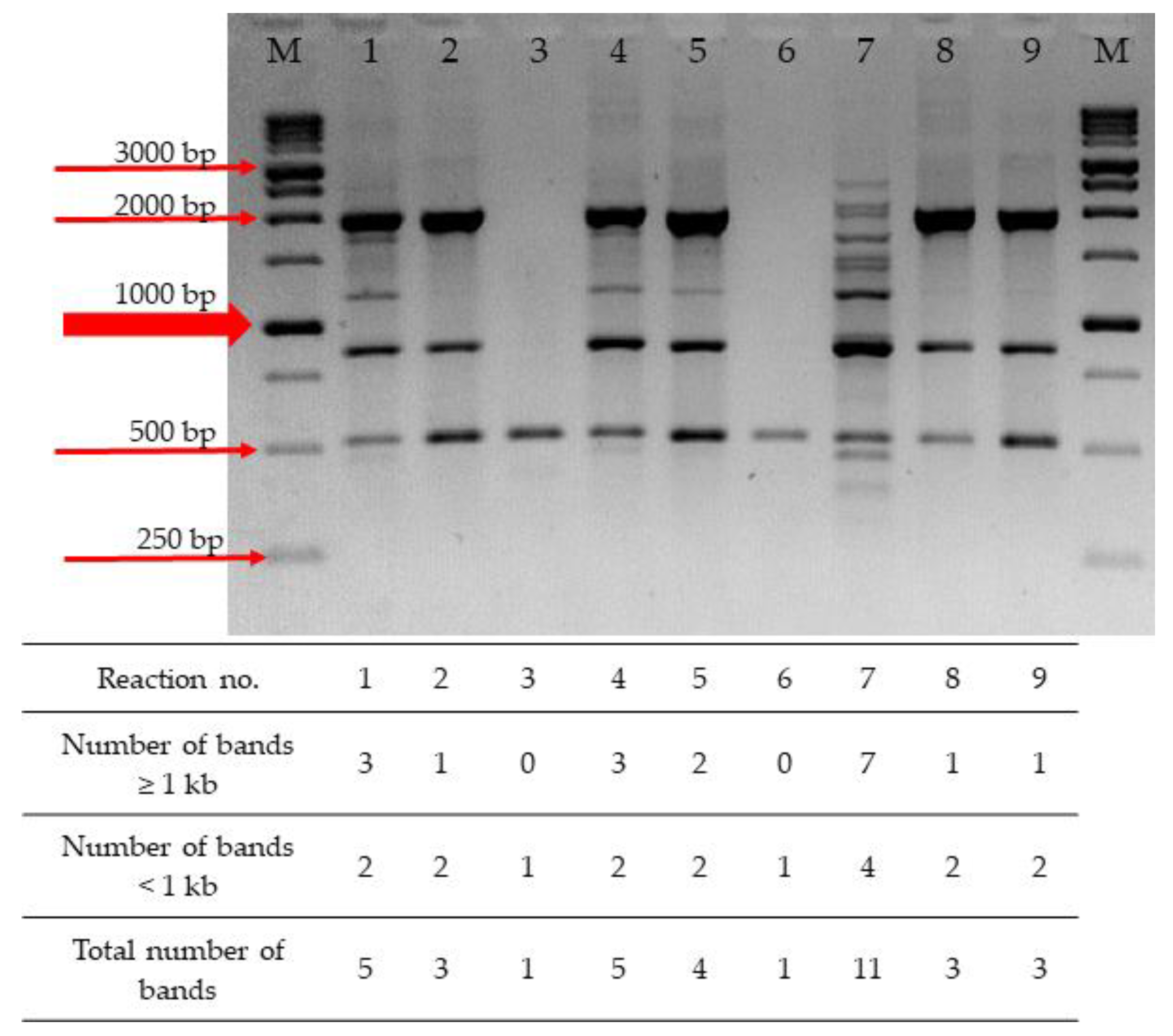
Pathogens | Free Full-Text | RAPD-PCR-Based Fingerprinting Method as a Tool for Epidemiological Analysis of Trueperella pyogenes Infections
View of A systematic method for DNA fragment amplification and sequencing based on DNA indexing technology. Protocol and technical considerations | Acta Biochimica Polonica
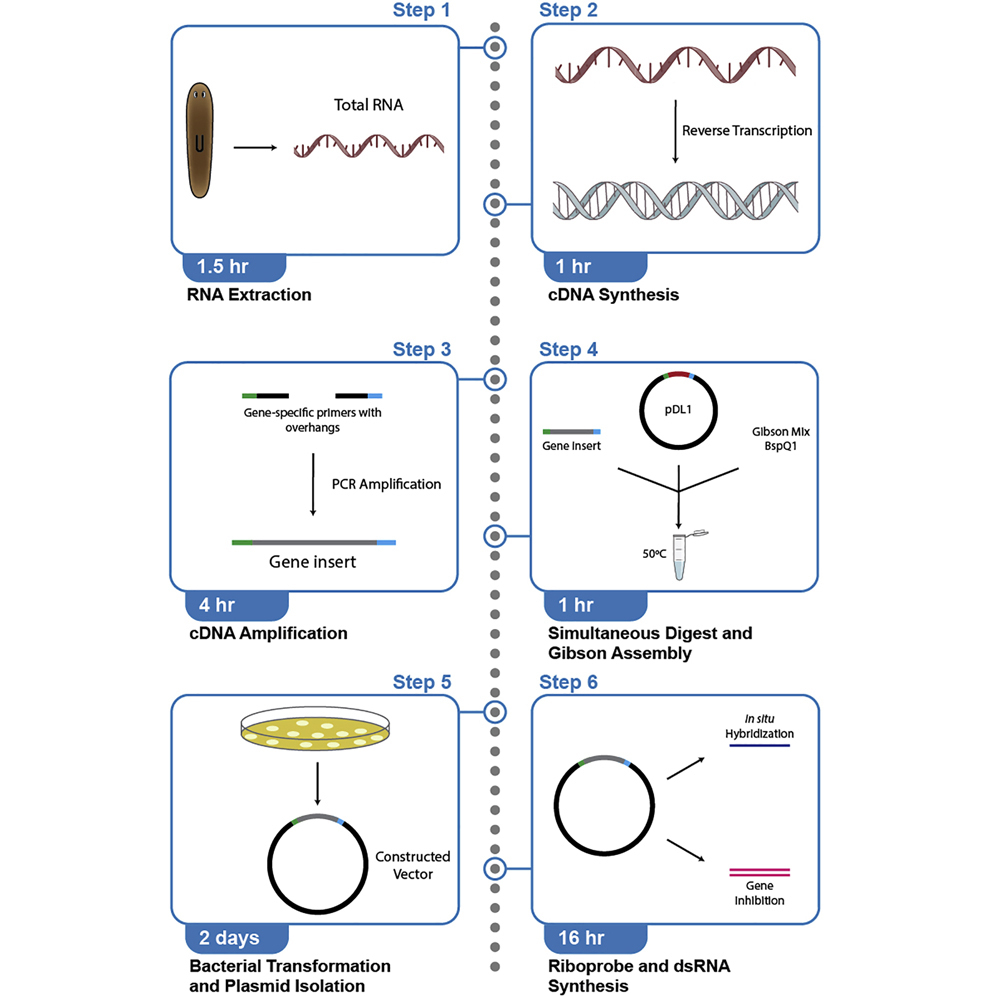
i>In situ</i> probe and inhibitory RNA synthesis using streamlined gene cloning with Gibson assembly

Rapid assessment of phytoplankton assemblages using Next Generation Sequencing – Barcode of Life database: a widely applicable toolkit to monitor biodiversity and harmful algal blooms (HABs) | bioRxiv
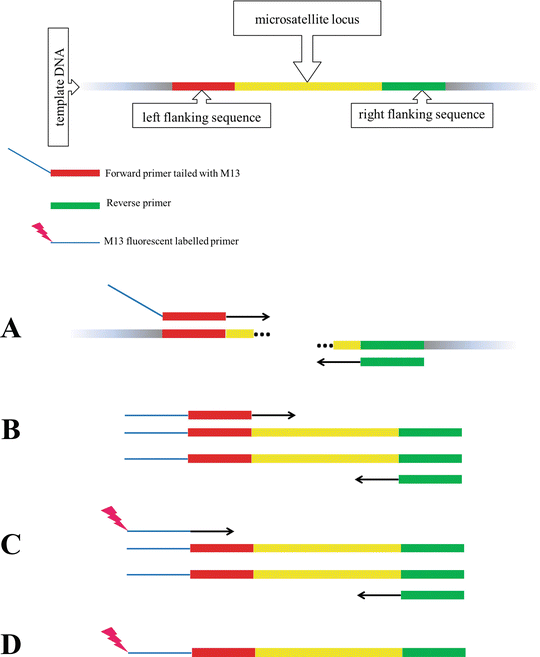
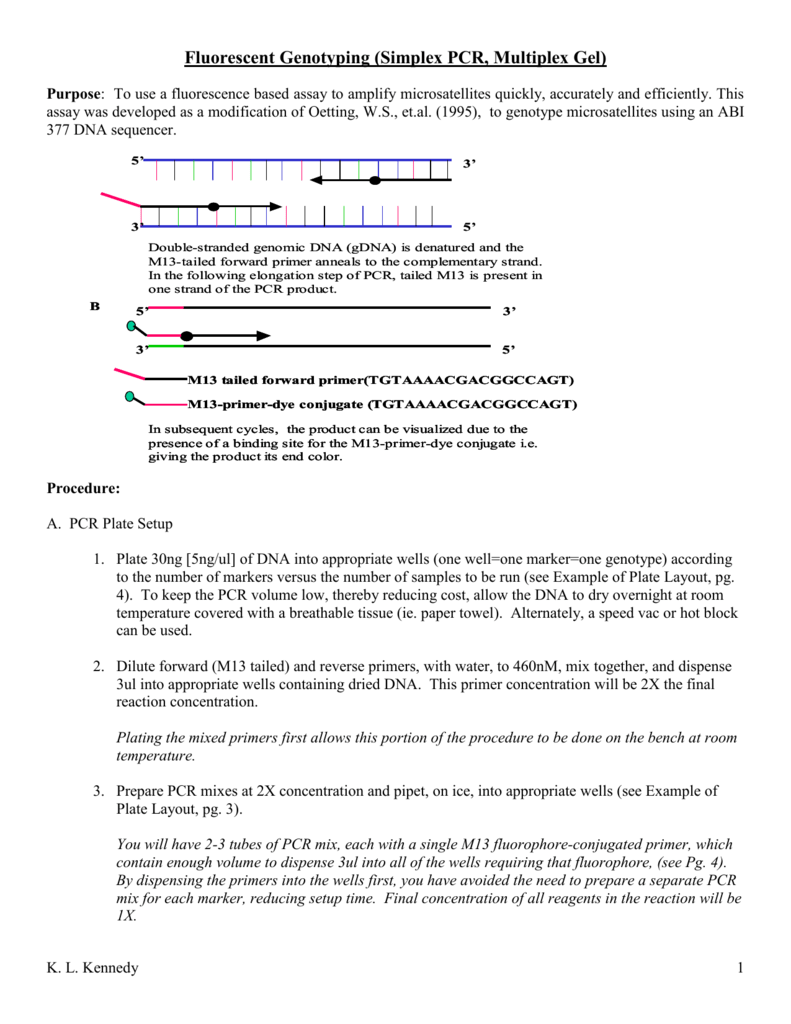
M13-Tailed Simple Sequence Repeat (SSR) Markers in Studies of Genetic Diversity and Population Structure of Common Oat Germplasm | SpringerLink

SciELO - Brasil - Adaptation of fluorescent technique for genotyping with new microsatellite markers in common bean Adaptation of fluorescent technique for genotyping with new microsatellite markers in common bean

Protocol to access unknown flanking DNA sequences using Wristwatch-PCR for genome-walking - ScienceDirect

A tailed PCR procedure for cost‐effective, two‐order multiplex sequencing of candidate genes in polyploid plants - Gholami - 2012 - Plant Biotechnology Journal - Wiley Online Library
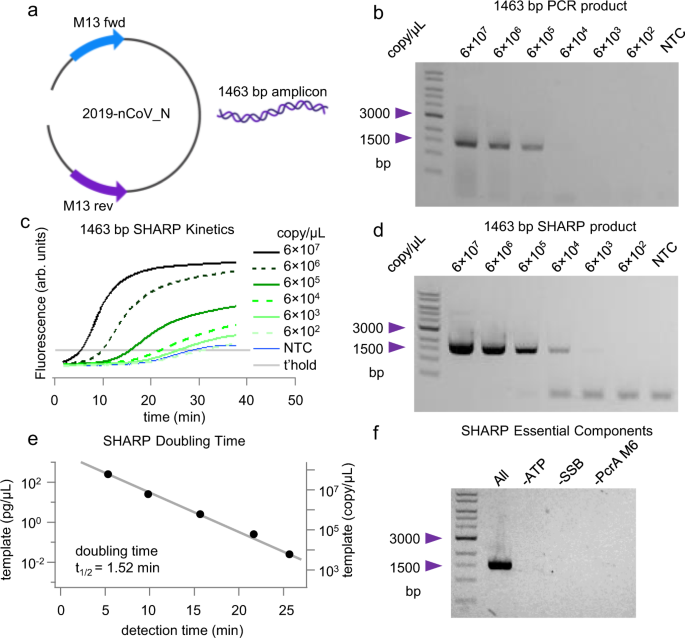
Engineered helicase replaces thermocycler in DNA amplification while retaining desired PCR characteristics | Nature Communications